
ore A ccurate Truong SK McCormick RF Morishige DT Mullet JE 2014 Resolution of genetic map expansion caused by excess heterozygosity in plant recombinant inbred populations G3 Bethesda 4 19631969 ID: 935282
Download Presentation The PPT/PDF document "Making Genetic M aps M" is the property of its rightful owner. Permission is granted to download and print the materials on this web site for personal, non-commercial use only, and to display it on your personal computer provided you do not modify the materials and that you retain all copyright notices contained in the materials. By downloading content from our website, you accept the terms of this agreement.
Slide1
Making
Genetic
Maps More Accurate
Truong SK, McCormick RF, Morishige DT, Mullet JE (2014) Resolution of genetic map expansion caused by excess heterozygosity in plant recombinant inbred populations. G3 (Bethesda) 4: 1963-1969
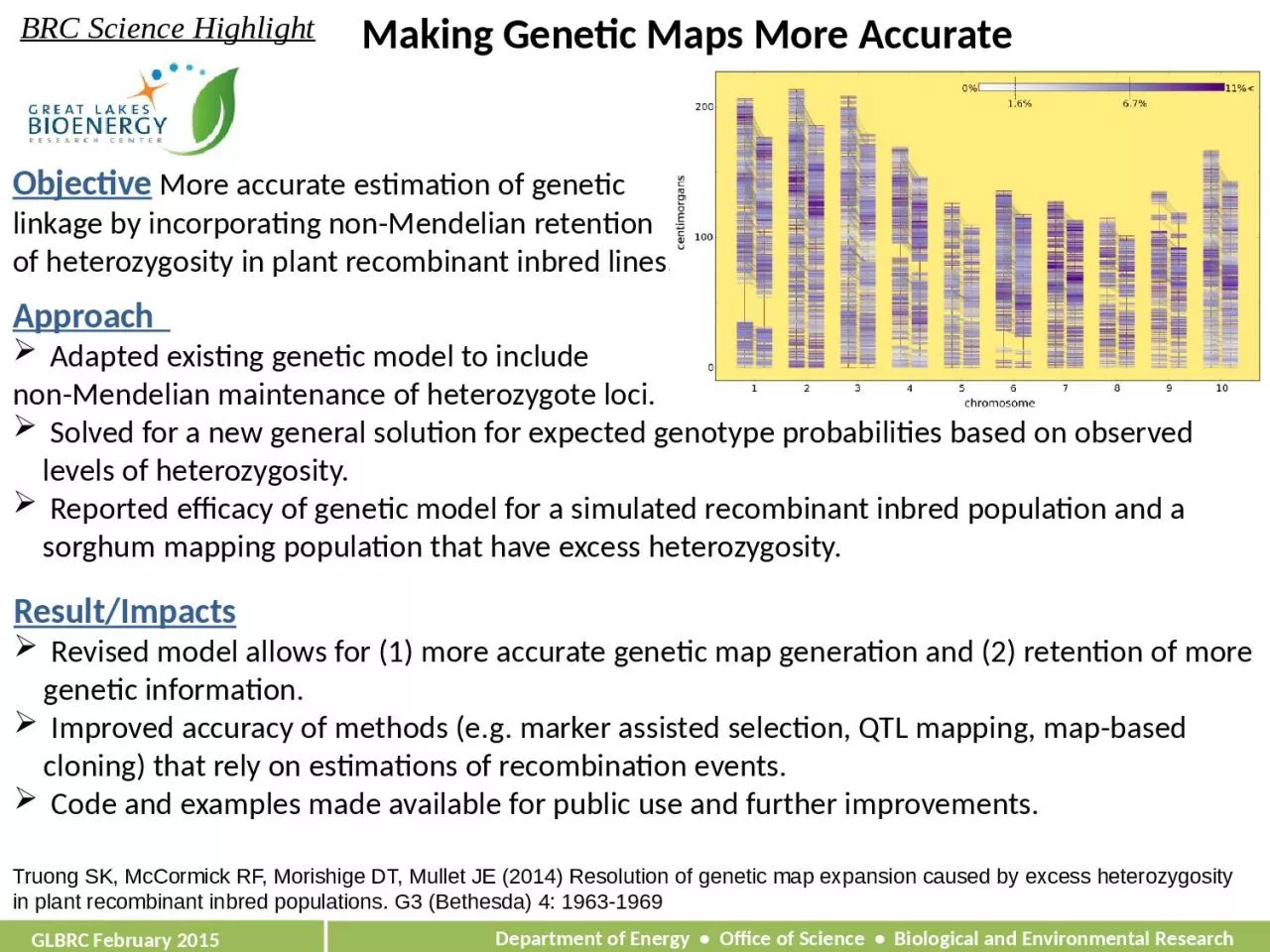
Objective More accurate estimation of genetic linkage by incorporating non-Mendelian retention of heterozygosity in plant recombinant inbred lines.
Approach Adapted existing genetic model to include non-Mendelian maintenance of heterozygote loci. Solved for a new general solution for expected genotype probabilities based on observed levels of heterozygosity. Reported efficacy of genetic model for a simulated recombinant inbred population and a sorghum mapping population that have excess heterozygosity.
Result/Impacts Revised model allows for (1) more accurate genetic map generation and (2) retention of more genetic information. Improved accuracy of methods (e.g. marker assisted selection, QTL mapping, map-based cloning) that rely on estimations of recombination events. Code and examples made available for public use and further improvements.
BRC Science Highlight
GLBRC
February 2015