
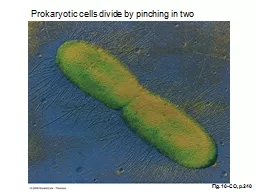
cells divide by pinching in two Learning Objectives 1 What Is the Flow of Genetic Information in the Cell 2 What Are the General Considerations in the Replication of DNA ID: 932744
Download Presentation The PPT/PDF document "Fig. 10-CO, p.240 Prokaryotic" is the property of its rightful owner. Permission is granted to download and print the materials on this web site for personal, non-commercial use only, and to display it on your personal computer provided you do not modify the materials and that you retain all copyright notices contained in the materials. By downloading content from our website, you accept the terms of this agreement.
Slide1
Fig. 10-CO, p.240
Prokaryotic
cells divide by pinching in two
Slide2Learning Objectives
1. What Is the Flow of Genetic Information in the Cell? 2. What Are the General Considerations in the
Replication of DNA? 3. How Does the DNA Polymerase Reaction Take Place? 4. Which Proteins Are Required for DNA Replication?
5. How Is DNA Replicated in Eukaryotes?
Slide3Th
Fig. 10-1, p.241
The
Central Dogma
The flow of
genetic information in biological systems.
Slide4It states that in all living organisms, the genetic information is stored in chromosomal DNA. The flow of genetic information occurs in one direction from DNA
RNA
Protein
Slide5Fig. 10-18, p.256
The eukaryotic cell cycle
Slide6DNA Replication
In this process the two polynucleotide chains of DNA are separated and each is copied in a complementary manner to produce daughter polynucleotide chains .Each
daughter DNA molecule will contain one polynucleotide comes from the parental DNA and the other chain is newly formed ( semiconservative replication)
Slide7Fig. 10-3, p.243
Experimental evidence for
semiconservative
replication.
Meselon
-Stahl Experiment
1958 ( In Prokaryotes )
Slide8Fig. 10-2, p.242
Slide9DNA
replication must occur in order to faithfully transmit genetic material to the progeny of any cell or organism. DNA replication takes place during S phase of the cell
cycle.Transcription is the process by which the information contained in a section of DNA is transferred to a newly assembled piece of messenger RNA (mRNA).Translation a process in which proteins can be synthesized using the information in mRNA as a template .
Slide10In
eukaryotic genome there is no similar linear relationship between genetic information carried in DNA and proteins expression, which was observed in prokaryotic systemOccasionally, genetic information flows from RNA to DNA (in reverse of normal transcription). This is known to occur in the case of retroviruses, such as HIV
.
Slide11DNA
replicationIn prokaryotes like E. coli , the replication starts at fixed point called the origin
and proceeds bidirectional along the chromosomal DNA (both directions) until the whole circular DNA is completely replicated. By then the replication process is terminated at certain point on DNA.
Slide12Fig. 10-4a, p.244
Slide13Fig. 10-5a, p.245
Slide14Fig. 10-5b, p.245
Slide15Mechanism of DNA replication in
ProkaryotesRelaxation of complex super structure of chromosomal DNAInside bacteria the circular double helix DNA is present in super helical form in which the DNA is further twisted in more circles. The relaxed double helix is called a secondary structure and the complex super helix called tertiary structure. The super helix is needed to pack the DNA in small space inside the cell but it is not suitable form for starting DNA replication. So, the first step in starting DNA replication is the relaxation of super coiled DNA by making small cut (nicking) with the enzyme Topoisomerase (
Gyrase).
Slide16Slide17DNA
replication involves 3 stages Initiation
, elongation and termination.1.Initiation:Initiation of DNA replication starts by the binding of the initiator factor protein
DnaA
at the initiation point in DNA, which recognizes a
repeat 9 A-T
base pair rich region. The binding of this protein helps in local unwinding double helix DNA at the initiation point by the enzyme
helicase
. After local unwinding the two single strands DNA are kept separated and not folded back again by
single strand binding proteins
.
Slide18Slide19Slide20Formation of bubble origin point
The separation of the two DNA strands expand the DNA region at the origin point to form a bubble shape area that help in the incorporation of
deoxynucleotides to synthesis new DNA strands.
Slide21Slide22II. Elongation of new DNA
At the bubble area, local unwinding and replication will grow in two opposite directions so that both polynucleotides are copied simultaneously .This polynucleotides growth form a Y-shaped replication fork at each direction site of DNA replication (2 replication forks).
Since, the DNA synthesis occurs in 5′ to 3′ direction, one strand called the leading strand, can be synthesized continuously while the other called the lagging strand, must be synthesized backward discontinuously in short fragments (Okazaki fragments) which are later joined to make one long piece.
Slide23Properties of prokaryotic DNA polymerase
enzymesThere are 3 different types of prokaryotic DNA polymerases ,called pol I,pol
II, pol III.Only pol I and III are involved in DNA replication while pol II function is limited to DNA repair of damaged DNA.The pol I enzyme has slower activity than pol III enzyme.Each DNA polymerase enzyme has two types of activities, polymerization and nuclease activities
.
Slide24Table 10-1, p.246
Slide25TH
t
Fig. 10-7, p.246
The dimer of
β
-subunits of DNA polymerase III bound to DNA
Slide261
) Polymerization activity
Add deoxynucleotids ( as monophosphate) from triphosphates deoxyncleotides , using energy liberated from the hydrolysis of pyrophosphates (the process is called polymerization).The monophosphate deoxynuleotides are connected by
phosphodiester
linkage in
5
3 direction
.
Slide27The building blocks for this process are 5'-ribonucleoside triphosphates, and pyrophosphate released as each
phosphodiester bonds made.
Slide28Slide29RNA primer
DNA polymerases cannot initiate synthesis of a complementary strand of DNA on a totally single-stranded template. Rather, they require an RNA primer that is,
a short, double-stranded region consisting of RNA base-paired to the DNA template, with a free OH-group on the 3'-end of the RNA strand .This OH group serves as the first acceptor of a nucleotide by action of DNA polymerase
. In de novo DNA synthesis, that free 3'-hydroxys provided by the short stretch of RNA, rather than DNA.
Slide30Primase
A specific RNA polymerase, called primase synthesizes the short stretches of RNA (approximately ten nucleotides long) that are complementary and antiparallel
to the DNA template. In the resulting hybrid duplex, the U in RNA pairs with A in DNA. These short RNA sequences are constantly being synthesized at the replication fork on the lagging strand, but only one RNA sequence at the origin of replication required on the leading strand.
Slide31Primosome
Prior to the beginning of RNA primer synthesis on the lagging strand, a
prepriming complex of several proteins assembled and binds to the single strand of DNA, displacing some of the single-stranded DNA-binding proteins. This protein complex, plus
primase
is called the
primosome
. It initiates Okazaki fragment formation by moving along the template for the lagging strand in
the 5’
3’ direction, periodically recognizing specific sequences of nucleotides that direct it to create an RNA primer
thats
synthesized in the 5’
3’ direction
(antiparallel
to the DNA template chain).
Slide322)
Exonuclease activity: Repeated cutting of single nucleotide from the terminal end of a polynucleotide
3-Exonuclease activity: Removal of polynucleotide sequence in 35 direction from the terminal end associated with pol III activity.5- Exonuclease
activity
: Removal of polynucleotide sequence in
5
3
direction from the terminal end, associated with pol I activity.
Slide33Fig. 10-11, p.250
DNA polymerase I proofreading removes nucleotides from the 3’ end of the growing DNA chain
.
Slide34Fig. 10-12a, p.252
The 5’
3’
exonuclease
activity of DNA polymerase I can remove up to 10 nucleotides in the 5’ direction downstream from a 3’-OH single –strand nick
Slide35Mechanism of DNA synthesis at each Replication
Fork1)-Leading strand
RNA primase enzyme creates a short primer RNA with free 3' end ( 10 RNA nucleotide sequence) . DNA polymerase III enzyme - uses a single parent strand of DNA as a template to add new nucleotides to the 3' OH end of initially incorporated RNA primer.
Slide36The addition is continuous according to the base pairing rule.
If a mismatch is accidentally incorporated, the polymerase is inhibited from further extension. Proofreading removes the mismatched nucleotide and extension continues. The mismatched nucleotides are remove by the exonuclease activities of DNA polymerase III.
Later , DNA polymerase I removes the RNA primers and replaces them by DNA pieces leaving a small gap of free 3' OH and 5’ OH ends to be sealed by DNA ligase .
Slide372)-Lagging strand
DNA polymerase is unable to work directly on the lagging strand because it lacks a free 3- OH end on the existing DNA strand.
The new strand is synthesized in short discontinuous segments, each segment consists of RNA primer formed by primase and replicated DNA piece (Okazaki fragments) about 100-200 DNA nucleotides in length formed by DNA polymerase III similarly to the leading strand except that the addition takes place in backward direction Later the polymerase I removes the RNA primers and replace them by DNA fragments
leaving gaps .
Ligase enzyme seals the gaps as described before..
Slide38Reiji Okazaki
(1930 –1975)was a pioneer Japanese
molecular biologist, known for his research on DNA replication and especially for describing the role of Okazaki fragments which he discovered working with his wife Tsuneko in 1966.
Okazaki was born in
Hiroshima
,
Japan
. He graduated in 1953 from
Nagoya University
, and worked as a professor there after 1963. He died of
leukemia
(due to
Atomic bombings of Hiroshima
) in 1975 at the age of 44; he had been
heavily
irradiated
in
Hiroshima
when the
first atomic bomb
was dropped. His wife,
Tsuneko
, won the
L'Oréal
-UNESCO Awards for Women in Science
in 2000 for her work
.
Slide39Slide40Slide41Slide42Final results of replication at the fork
level
Slide43III.Termination
:Termination
requires that the progress of the DNA replication fork must stop at a specific locus on DNA. This process involves the interaction between two components:(1) A termination site sequence in the DNA(2) A protein which binds to this sequence to physically stop DNA replication.In bacteria this protein is named the DNA replication terminus site-binding protein.Because bacteria have circular chromosomes, termination of replication occurs when the two replication forks meet each other on the opposite end of the parental chromosome.
In E. coli the chromosomal DNA replication takes about 40 minutes to replicate the 4000 kb size of DNA. Therefore each fork replicates 2000 kb in 40 min. or ~ 50 kb/min or ~1000 bases/sec.
Slide44p.251a
Why Does DNA Contain Thymine & Not Uracil ?
Slide45The incorporation of thymine instead of uracil helps ensure that the DNA is replicated faithfully.
p.251b
Slide46Slide47Fig. 10-10, p.249
General features of a replication fork
Slide48Table 10-3, p.250
Slide49END Part I
Slide50Eukaryotic DNA replication
Slide51Eukaryotic genome is more complex than prokaryotic genome. For example human genome is composed of
30,000-40,000 genes, and each gene is segmented into two types of DNA pieces, exons and introns ,Each gene has on average between
5 and 8 exons, 8000 base-pairs.
Slide52Exon:
DNA segment which after transcription to RNA codes directly to peptide units of a polypeptide, i.e which `is expressed in protein' (200 base-pairs on average, in human genome
)Intron: DNA segment which is not directly expressed for protein, involved in regulation, splicing and other unknown functions each 2000 bp's on average, in human genome.
Slide53Cell
cycle in eukaryotes
Actively dividing eukaryote cells pass through a series of stages known collectively as the cell cycle: involving interphase stage between each two mitosis. The interphase stage itself includes 3 phases represented by two gap phases (G1 and G2); and S (for synthesis) phase, in which the genetic material is duplicated. The two gaps are preparative periods for cell division and an opportunity for the cell to make decision whether to go in to division or not. In the M phase a nuclear division takes place before it is followed by cell division that occurs in cytokinesis
Slide54Slide55In the S phase
. DNA synthesis replicates the genetic material and each chromosome becomes having two sister chromatids. Therefore, only in this period of the interphase that DNA replication occurs which is a necessary step before the cell decides to go in to division(without DNA replication there is no cell division).In human the S period is carried out for about
8 hours in an average total cell cycle period of around 20 hours.
Slide56Eukaryotic DNA synthesis is similar to synthesis in prokaryotes, except for some complexity. In eukaryotic cells:
there is more DNA than prokaryotic cellsthe chromosomes are linear
the DNA complexes with proteins
Slide57Eukaryotic
replication initiated at many points
Because the eukaryotic genome is so large (about 100 times the size of bacterial DNA), it would take days to replicate the whole length of eukaryotic chromosome using the same single initiation point as in prokaryotes.Therefore, many initiation points ( about 10,000 in human) are found in each eukaryotic chromosome instead of one, with replication forks moving in opposite directions away from each initiation point until they meet in the middle between each two initiation points. The initiation point is called a
replicons
which do not need specific termination sequences.
Slide58Fig. 10-4b, p.244
Slide59Not all replicons are activated simultaneously. Rather, clusters of
20-80 adjacent replicons are activated throughout S phase until the whole chromosome is completely replicated.
Slide60The rate of eukaryotic DNA replication is much slower than
E. coli, with only 100-200 nucleotides bases/sec are replicated in eukaryotic Okazaki fragments. However, the majority of replication forks results in the whole genome being replicated in only about 8 hours. Histones for packaging the DNA are synthesized simultaneously with DNA replication to bind the new DNA.
Slide61Enzymes
involved in eukaryotic DNA replication
1.DNA Polymerase αInitiation the synthesis of RNA primer (about 20-30 ribonucleotides) then adds DNA to the RNA primers It has low processivity
(efficiency) of DNA synthesis and has no
3
5
exonuclease
activity .
2.DNA Polymerase
δ
The
principal DNA polymerase in eukaryotic DNA replication which has
3
5
exonuclease
activity.
When
it complexes with
PCNA
(Proliferating Cell Nuclear Antigen) it becomes highly
processive
.
Slide62Additional
Proteins Involved in Eukaryotic DNA Synthesis
DNA helicase: the enzyme which carries out partial unwinding of double helix DNA at the initiation point before the starting of DNA replicationPCNA (Proliferating Cell Nuclear Antigen) Provides high processivity to DNA Polymerase δ
RPA
(Replication Protein A)
ssDNA
-binding protein
that facilitates the unwinding of the helix to create two replication forks.
RFC
(Replication Factor C) binds PCNA at the end of the primer
FEN1/RTH1
(flap endonuclease 1/RAD two homologue 1)
exonuclease
complex
Slide63Leading
strand synthesis
1) Starts with the primase activity of DNA Pol α to put down RNA primer in 5’ 3’-direction2) The same enzyme adds a piece of DNA to the primer3) RFC binds PCNA at the end of the primer
4) PCNA displaces DNA Pol
α
.
5) DNA polymerase
δ
binds to PCNA at the 3’ ends of the growing strand to carry out polymerase switching to highly
processive
DNA synthesis activity.
The RFC mediates the polymerase switching by helping in the
a) Assembly of PCNA
b) Removal of DNA Pol
α
c) Addition of DNA Pol
δ
Slide64Lagging strand synthesis
1) Starts off the same way as leading strand synthesis
2) RNA primers synthesized by DNA polymerase α every 50 nucleotides and consist of 20-30 nucleotides RNA3) DNA polymerase δ switching as before to extend the RNA
primers
and generating Okazaki fragments
4) When the DNA Pol δ polymerizes the RNA primer of the
downstream
Okazaki fragment,
RNase
H1 removes all but the
last
RNA nucleotide of the RNA primer
5) The FEN1/RTH1
exonuclease
complex removes the last RNA
nucleotide
6) DNA Pol δ fills in the gap as the RNA primer is being
removed
7) DNA ligase joins the Okazaki fragment to the growing strand
Slide65Telomeres
problem during human DNA replication
Telomeres present at the ends of linear chromosomal DNA and consist of long area of short repeating sequences TTGGGG ->->->->- to protect the integrity and stability of human chromosomes. During DNA synthesis these chromosome ends cannot be replicated with DNA polymerase.
Slide66This sequence of TTAGGG is repeated approximately
2,500 times in humans. In humans, average telomere length declines from about 11
kilobases at birth to less than 4 kilobases in old age, with average rate of decline being greater in men than in women.
Slide67Telomeres are found at the termini of chromosomes. The end of a telomere inserts back into the main body of the telomere to form a T-loop
Slide68Since DNA can only be synthesized at the 3'-end of a pre-exiting DNA or RNA chain,
there is no available mechanism for achieving DNA synthesis all of the ways to the end of the lagging strand. Once the primer in the last Okasaki
fragment is removed by a 5' to 3' exonuclease it is not possible to replace it with DNA. This is because the 5 - ends of the lagging strands does not have enough space to put a new primer with free 3'-hydroxyl group and therefore it is not copied completely.
Slide69After
maturation of Okazaki fragments, there is a primer gap
Slide70This problem does not occur with the leading strand which can undergo complete
replication round. If this phenomenon is repeated over many rounds of replication, some of the chromosomes will gradually develop major shortening in their ends.
Slide71Slide72Slide73Correction of the chromosome ends by the enzyme Telomerase
Telomerase
enzyme is a ribonucleo protein complex containing RNA-dependent DNA polymerase activity and 450-nucleotide RNA. It can act as a reverse transcriptase
enzyme
by using its own repetitive RNA sequence (AAAACCCC ) as a template to add
a repeat
complementary sequence of TTAGGG to the 3- OH end of leading strand
in telomeres
of human DNA. This addition step by telomerase is repeated several times
until an
extend 3- end of the DNA is formed. In this case the role of telomerase enzyme
is ended
leaving gap in the 5- phosphate end of the opposite lagging DNA strand .
This
gap will
be filled later by
primase
(adding short RNA primer ) and combined actions of
DNA polymerase
and ligase activities
..
Slide74Slide75Slide76Some
somatic cells lack telomerase activity and therefore, their telomeres get shorter with each cell division (About 50 bases are lost from each telomere every time a cell divides) which may end up with cell death. To the contrary cancer cells
may have high activity of telomerase enzyme that increases their survivals,
Slide77Telomerase
: Terminates the process of DNA replication only at the telomere ends of chromosomal DNA by adding many repeat units that can not be recognized by the replication complex.
Slide78p.259a
Slide79p.259b
Slide80p.258a
Slide81p.258b
Telomere
replication
(asterisks indicate sequences at the 3’ end that cannot be copied by conventional DNA replication
Slide82Model for initiation of the DNA replication cycle in eukaryotes
ORC is present at the replicators throughout the cell cycle.
The pre-replication complex (pre-RC) is assembled through sequential addition of the RAP (replication activator protein) & RLFs (replication licensing factors ) during a window of opportunity defined by the state of cyclin
-CDKs.
Phosphorylation of the
RAP,ORC,and
RLFs triggers replication.
After initiation, a post-RC state is established, and the RAP & RLFs are degraded.
Slide83Fig. 10-19, p.257
Model for initiation of the DNA replication cycle in eukaryotes
Slide84Fig. 10-21, p.261
The basics of the eukaryotic replication fork
Slide85Table 10-4, p.257
Slide86Table 10-5, p.260
Slide87END
Part II