
K S Myers et al Rewired cellular signaling coordinates sugar and hypoxic responses for anaerobic xylose fermentation in yeast PLoS Genetics 15 e1008037 2019 101371journalpgen1008037 ID: 1030528
Download Presentation The PPT/PDF document "Xylose fermentation by yeast requires re..." is the property of its rightful owner. Permission is granted to download and print the materials on this web site for personal, non-commercial use only, and to display it on your personal computer provided you do not modify the materials and that you retain all copyright notices contained in the materials. By downloading content from our website, you accept the terms of this agreement.
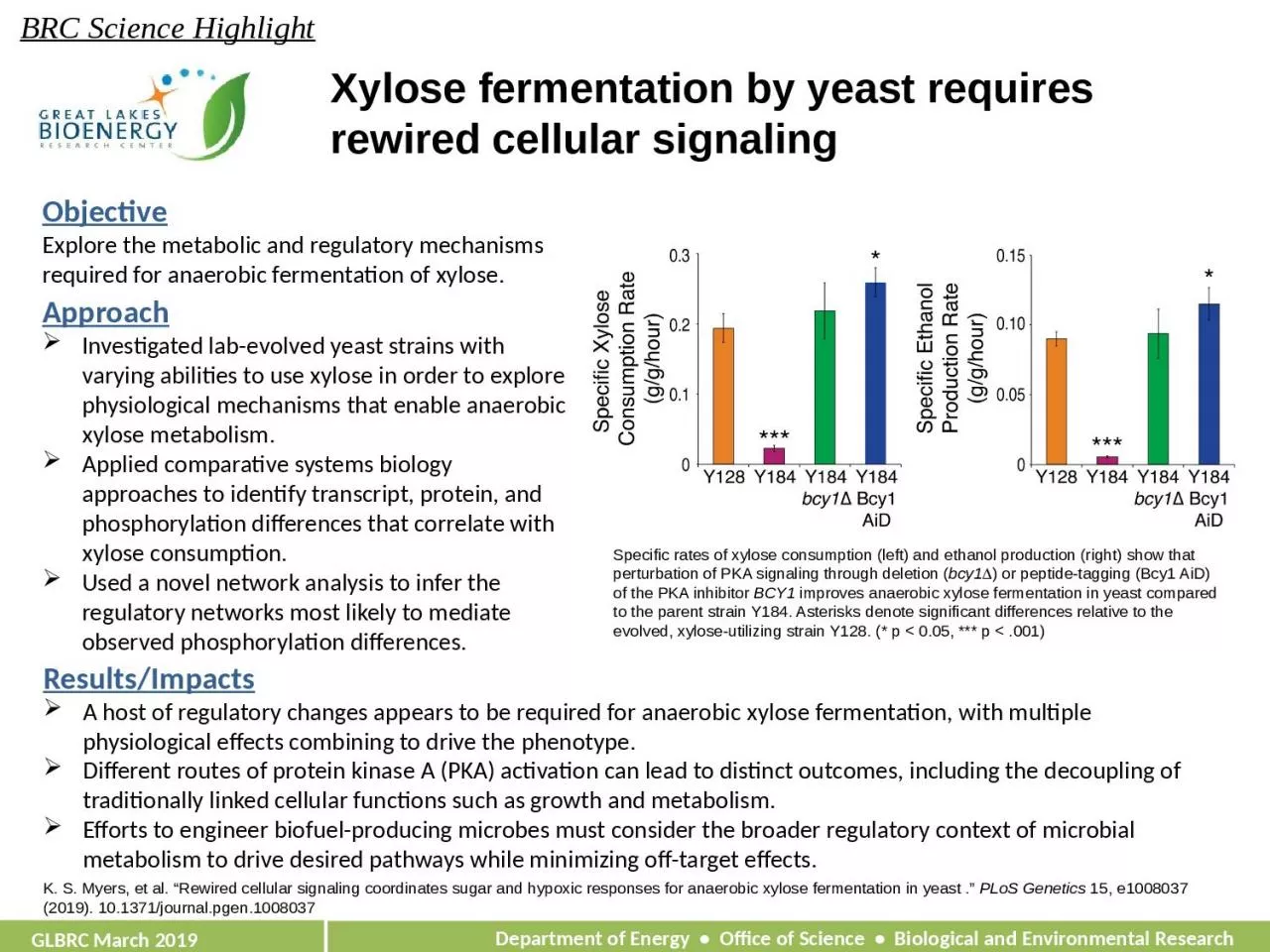
1. Xylose fermentation by yeast requires rewired cellular signalingK. S. Myers, et al. “Rewired cellular signaling coordinates sugar and hypoxic responses for anaerobic xylose fermentation in yeast .” PLoS Genetics 15, e1008037 (2019). 10.1371/journal.pgen.1008037Objective Explore the metabolic and regulatory mechanisms required for anaerobic fermentation of xylose.Results/ImpactsA host of regulatory changes appears to be required for anaerobic xylose fermentation, with multiple physiological effects combining to drive the phenotype.Different routes of protein kinase A (PKA) activation can lead to distinct outcomes, including the decoupling of traditionally linked cellular functions such as growth and metabolism. Efforts to engineer biofuel-producing microbes must consider the broader regulatory context of microbial metabolism to drive desired pathways while minimizing off-target effects.BRC Science Highlight GLBRC March 2019Specific rates of xylose consumption (left) and ethanol production (right) show that perturbation of PKA signaling through deletion (bcy1D) or peptide-tagging (Bcy1 AiD) of the PKA inhibitor BCY1 improves anaerobic xylose fermentation in yeast compared to the parent strain Y184. Asterisks denote significant differences relative to the evolved, xylose-utilizing strain Y128. (* p < 0.05, *** p < .001)ApproachInvestigated lab-evolved yeast strains with varying abilities to use xylose in order to explore physiological mechanisms that enable anaerobic xylose metabolism.Applied comparative systems biology approaches to identify transcript, protein, and phosphorylation differences that correlate with xylose consumption.Used a novel network analysis to infer the regulatory networks most likely to mediate observed phosphorylation differences.