
Probability Introduction to Biostatistics and Bioinformatics Sequence Alignment Concepts This Lecture Sequence Alignment Stuart M Brown PhD Center for Health Informatics and Bioinformatics ID: 308018
Download Presentation The PPT/PDF document "Previous Lecture:" is the property of its rightful owner. Permission is granted to download and print the materials on this web site for personal, non-commercial use only, and to display it on your personal computer provided you do not modify the materials and that you retain all copyright notices contained in the materials. By downloading content from our website, you accept the terms of this agreement.
Slide1
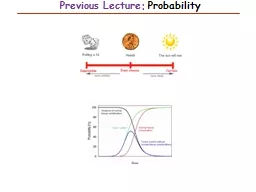
Previous Lecture: ProbabilitySlide2
Introduction to Biostatistics and Bioinformatics
Sequence Alignment Concepts
This LectureSlide3
Sequence Alignment
Stuart M. Brown, Ph.D.
Center for Health Informatics and Bioinformatics
NYU School of Medicine
Slides/images/text/examples borrowed liberally from:
Torgeir R. Hvidsten,
Michael Schatz,
Bill Pearson,
Fourie Joubert, others …Slide4
Learning Objectives
Identity, similarity, homology
Analyze sequence similarity by dotplots
window/stringency
Alignment of text strings by edit distance
Scoring of aligned amino acids
Gap penalties
Global vs. local alignment
Dynamic Programming (Smith Waterman)
FASTA methodSlide5
Why Compare Sequences?
Identify sequences found in lab experiments
What is this thing I just found?Compare new genes to known ones
Compare genes from different species
information about evolution
Guess functions for entire genomes full of new gene sequences
Map sequence reads to a Reference Genome
(ChIP-seq, RNA-seq, etc.)Slide6
Are there other sequences like this one?
1) Huge public databases - GenBank, Swissprot, etc.2) “Sequence comparison is the most powerful and reliable method to determine evolutionary relationships between genes” -R. Pearson
3) Similarity searching is based on alignment
4)
BLAST
and
FASTA
provide rapid similarity searching
a. rapid = approximate (heuristic)
b. false + and - scoresSlide7
Similarity ≠ Homology
1) 25% similarity ≥ 100 AAs is strong evidence for homology
2) Homology is an evolutionary statement which means “descent from a common ancestor” common 3D structure
usually common function
homology is all or nothing, you cannot say "50% homologous"Slide8
Similarity is Based on Dot Plots
1) two sequences on vertical and horizontal axes of graph
2) put dots wherever there is a match
3) diagonal line is region of identity
(local alignment)
4) apply a window filter - look at a group of bases, must meet % identity to get a dot Slide9
Simple Dot PlotSlide10
PTHPLASKTQILPEDLASEDLTI
PTHPLAGERAIGLARLAEEDFGM
Score = 7
PTHPLASKTQILPEDLASEDLTI
PTHPLAGERAIGLARLAEEDFGM
Score = 11
Window = 12
Stringency = 9
Filtering
PTHPLASKTQILPEDLASEDLTI
PTHPLAGERAIGLARLAEEDFGM
Score = 11
Window / StringencySlide11
Dot plot filtered with 4 base window and 75% identitySlide12
Dot plot of real dataSlide13
Hemoglobin
-chain
Hemoglobin
-
chain
Dotplot
(Window = 130 / Stringency = 9)Slide14
Dotplot
(Window = 18 / Stringency = 10)
Hemoglobin
-
chain
Hemoglobin
-chainSlide15
Manually line them up and count?an alignment program can do it for you
or a just use a text editor
Dot Plot
shows regions of similarity as diagonals
GATG
C
C
A
T
A
G
A
G
C
T
G
T
A
G
T
C
GT
A
CCC
T <
—
—
>
C
T
A
G
A
G
A
G
C
-GT
AGT
CAGAGTGTCTTTGAGTTCC
How to Align Sequences?Slide16
Percent Sequence Identity
The extent to which two nucleotide or amino acid sequences are invariant
A C C T G A G – A G
A C G T G – G C A G
70% identical
mismatch
indelSlide17
Hamming DistanceThe minimum number of base changes that will convert one (ungapped) sequence into another
The Hamming distance is named after Richard
Hamming, who introduced it in his fundamental paper on Hamming codes:
“Error detecting and error correcting codes” (1950) Bell System Technical Journal 29 (2): 147–160.
Python function hamming_distance
def
hamming_distance(s1
,
s2):
#Return the Hamming distance between equal-length sequences
if
len
(s1)
!=
len
(s2):
raise
ValueError
(
"Undefined for sequences of unequal length"
)
return
sum
(ch1
!=
ch2
for
ch1
,
ch2
in
zip
(s1, s2))Slide18
Hamming Dist can be unrealistic
v: A
T
A
T
A
T
A
T
w: T
A
T
A
T
A
T
A
Hamming Dist
= 8 (no gaps, no shifts)
Levenshtein (1966) introduced
edit distance
v = _
A
T
A
T
A
T
A
T
w = T
A
T
A
T
A
TA_
edit distance
:
d
(
v
,
w
) = 2 Levenshtein, Vladimir I. (February 1966). "Binary codes capable of correcting deletions, insertions, and reversals".
Soviet Physics Doklady 10 (8): 707–710.Slide19
Affine Gap PenaltiesSlide20
Gap PenalitesWith unlimited gaps (no penalty), unrelated sequences can align (especially DNA)
Gap should cost much more than a mismatchMulti-base gap should cost only a little bit more than a single base gap
Adding an additional gap near another gap should cost more (not implemented in most algorithms)Score for a gap of length
x is: -(
p
+
σ
x)
p
is
gap open
penalty
σ
is
gap extend
penaltySlide21
Global
vs
Local similarity
Global
similarity uses complete aligned sequences - total % matches
- Needleman
&
Wunch
algorithm
2)
Local
similarity looks for best internal matching region between 2 sequences
- find a diagonal region on the
dotplot
Smith
-Waterman
algorithm
BLAST
and
FASTA
3) dynamic programming
optimal computer solution, not approximateSlide22
Global vs. Local AlignmentsSlide23Slide24
[Essentially finding the diagonals on the dotplot]
Basic principles of dynamic programming
- Creation of an alignment path matrix
- Stepwise calculation of score values
- Backtracking (evaluation of the optimal path)
Smith-Waterman MethodSlide25
Creation of an alignment path matrix
Idea: Build up an optimal alignment using previous solutions for optimal alignments of smaller subsequences
Construct matrix
F
indexed by
i
and
j
(one index for each sequence)
F
(
i,j
) is the score of the best alignment between the initial segment
x
1...
i
of
x
up to
x
i
and the initial segment
y
1...
j
of
y
up to
y
j
Build
F
(
i,j
) recursively beginning with
F(0,0) = 0Slide26
Michael SchatzSlide27
If F
(i-1,j-1),
F(i-1,j
) and
F
(
i
,
j
-1) are known we can calculate
F
(
i
,
j
)
Three possibilities:
x
i
and
y
j
are aligned,
F
(
i
,
j
) =
F
(
i
-1,
j-1) + s(x
i ,yj)
xi is aligned to a gap, F
(i,j) =
F(i-1,j
) - d y
j is aligned to a gap, F(
i,j) = F(
i,j-1) - d
The best score up to (i,j) will be the smallest of the three options
Creation of an alignment path matrixSlide28
Michael Schatz
Michael SchatzSlide29
Michael SchatzSlide30
Smith-Waterman is OPTIMAL but computationally slow
SW search requires computing of matrix of scores at every possible alignment position with every possible gap.
Compute task increases with the product of the lengths of two sequence to be compared (N2
)
Difficult for comparison of one small sequence to a much larger one, very difficult for two large sequences, essentially impossible to search very large databases. Slide31
Scoring Similarity
1) Can only score aligned sequences
2) DNA is usually scored as identical or not3) modified scoring for gaps - single vs. multiple base gaps (gap extension)
4) Protein AAs have varying degrees of similarity
a. # of mutations to convert one to another
b. chemical similarity
c. observed mutation frequencies
5) PAM matrix calculated from observed mutations in protein familiesSlide32
Amino acids have different biochemical and physical properties
that influence their relative replaceability in evolution.
C
P
G
G
A
V
I
L
M
F
Y
W
H
K
R
E
Q
D
N
S
T
C
SH
S+S
positive
charged
polar
aliphatic
aromatic
small
tiny
hydrophobic
Protein
Scoring SystemsSlide33
The PAM 250 scoring matrixSlide34
PAM Vs. BLOSUM
PAM100 = BLOSUM90
PAM120 = BLOSUM80
PAM160 = BLOSUM60
PAM200 = BLOSUM52
PAM250 = BLOSUM45
More distant sequences
BLOSUM62 for general use
BLOSUM80 for close relations
BLOSUM45 for distant relations
PAM120 for general use
PAM60 for close relations
PAM250 for distant relationsSlide35
Search with Protein, not DNA Sequences
1) 4 DNA bases vs. 20 amino acids - less chance similarity
2) can have varying degrees of similarity between different AAs
- # of mutations, chemical similarity, PAM matrix
3) protein databanks are
much
smaller than DNA databanksSlide36
FASTA
1) A faster method to find similarities and make alignments – capable of searching many sequences (an entire database)
2) Only searches near the diagonal of the alignment matrix
3) Produces a statistic for each alignment (more on this in the next lecture)Slide37
FASTA
1) Derived from logic of the dot plot
compute best diagonals from all frames of alignment2) Word method looks for exact matches between words in query and test sequence
hash tables (fast computer technique)
Only matches exactly identical words
DNA words are usually 6 bases
protein words are 1 or 2 amino acids
only searches for diagonals in region of word
matches = faster searchingSlide38
FASTA Format
simple format used by almost all programs>header line with a [return] at end
Sequence (no specific requirements for line length, characters, etc)
>URO1 uro1.seq Length: 2018 November 9, 2000 11:50 Type: N Check: 3854 ..
CGCAGAAAGAGGAGGCGCTTGCCTTCAGCTTGTGGGAAATCCCGAAGATGGCCAAAGACA
ACTCAACTGTTCGTTGCTTCCAGGGCCTGCTGATTTTTGGAAATGTGATTATTGGTTGTT
GCGGCATTGCCCTGACTGCGGAGTGCATCTTCTTTGTATCTGACCAACACAGCCTCTACC
CACTGCTTGAAGCCACCGACAACGATGACATCTATGGGGCTGCCTGGATCGGCATATTTG
TGGGCATCTGCCTCTTCTGCCTGTCTGTTCTAGGCATTGTAGGCATCATGAAGTCCAGCA
GGAAAATTCTTCTGGCGTATTTCATTCTGATGTTTATAGTATATGCCTTTGAAGTGGCAT
CTTGTATCACAGCAGCAACACAACAAGACTTTTTCACACCCAACCTCTTCCTGAAGCAGA
TGCTAGAGAGGTACCAAAACAACAGCCCTCCAAACAATGATGACCAGTGGAAAAACAATG
GAGTCACCAAAACCTGGGACAGGCTCATGCTCCAGGACAATTGCTGTGGCGTAAATGGTC
CATCAGACTGGCAAAAATACACATCTGCCTTCCGGACTGAGAATAATGATGCTGACTATC
CCTGGCCTCGTCAATGCTGTGTTATGAACAATCTTAAAGAACCTCTCAACCTGGAGGCTTSlide39
FASTA AlgorithmSlide40
Makes Longest Diagonal
3) after all diagonals found, tries to join diagonals by adding gaps
4) computes alignments in regions of best diagonalsSlide41
FASTA AlignmentsSlide42
FASTA Results - Alignment
SCORES Init1: 1515 Initn: 1565 Opt: 1687 z-score: 1158.1 E(): 2.3e-58
>>GB_IN3:DMU09374 (2038 nt)
initn: 1565 init1: 1515 opt: 1687 Z-score: 1158.1 expect(): 2.3e-58
66.2% identity in 875 nt overlap
(83-957:151-1022)
60 70 80 90 100 110
u39412.gb_pr CCCTTTGTGGCCGCCATGGACAATTCCGGGAAGGAAGCGGAGGCGATGGCGCTGTTGGCC
|| ||| | ||||| | ||| |||||
DMU09374 AGGCGGACATAAATCCTCGACATGGGTGACAACGAACAGAAGGCGCTCCAACTGATGGCC
130 140 150 160 170 180
120 130 140 150 160 170
u39412.gb_pr GAGGCGGAGCGCAAAGTGAAGAACTCGCAGTCCTTCTTCTCTGGCCTCTTTGGAGGCTCA
||||||||| || ||| | | || ||| | || || ||||| ||
DMU09374 GAGGCGGAGAAGAAGTTGACCCAGCAGAAGGGCTTTCTGGGATCGCTGTTCGGAGGGTCC
190 200 210 220 230 240
180 190 200 210 220 230
u39412.gb_pr TCCAAAATAGAGGAAGCATGCGAAATCTACGCCAGAGCAGCAAACATGTTCAAAATGGCC
||| | ||||| || ||| |||| | || | |||||||| || ||| ||
DMU09374 AACAAGGTGGAGGACGCCATCGAGTGCTACCAGCGGGCGGGCAACATGTTTAAGATGTCC
250 260 270 280 290 300
240 250 260 270 280 290
u39412.gb_pr AAAAACTGGAGTGCTGCTGGAAACGCGTTCTGCCAGGCTGCACAGCTGCACCTGCAGCTC
|||||||||| ||||| | |||||| |||| ||| || ||| || |
DMU09374 AAAAACTGGACAAAGGCTGGGGAGTGCTTCTGCGAGGCGGCAACTCTACACGCGCGGGCT
310 320 330 340 350 360Slide43
Summary
Identity, similarity, homology
Analyze sequence similarity by dotplots
window/stringency
Alignment of text strings by edit distance
Scoring of aligned amino acids
Gap penalties
Global vs. local alignment
Dynamic Programming (
Smith Waterman
)
FASTA methodSlide44
Next Lecture: Searching Sequence Databases