
BRCA2 Variants by SiteDirected Mutagenesis Meredith Ellington Sarah Brnich 12 Jonathan S Berg 1 1 Department of Genetics UNC School of Medicine 2 Curriculum in Genetics amp Molecular Biology UNC School of Medicine ID: 913077
Download Presentation The PPT/PDF document "Results Generation of Pathogenic and Ben..." is the property of its rightful owner. Permission is granted to download and print the materials on this web site for personal, non-commercial use only, and to display it on your personal computer provided you do not modify the materials and that you retain all copyright notices contained in the materials. By downloading content from our website, you accept the terms of this agreement.
Slide1
Results
Generation of Pathogenic and Benign
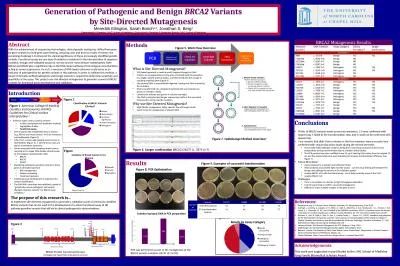
BRCA2 Variantsby Site-Directed MutagenesisMeredith Ellington, Sarah Brnich1,2, Jonathan S. Berg11Department of Genetics, UNC School of Medicine; 2Curriculum in Genetics & Molecular Biology, UNC School of Medicine
AbstractWith the advancement of sequencing technologies, clinical genetic testing has shifted from gene-by-gene analysis to multi-gene panel testing, reducing costs and time to results. However, the remaining challenge is to interpret the clinical significance of these increasingly identified genetic variants. Functional assays are one type of evidence considered in the interpretation of sequence variation, though well-validated assays do not yet exist for most disease-related genes. Both BRCA2 and PALB2 play a significant role in the DNA repair pathway of homologous recombination, acting as tumor suppressors. As such, a measure of DNA repair outcomes could serve as an indicator of pathogenicity for genetic variants in this pathway. In order to validate this method, a panel of clinically verified pathogenic and benign variants is required to determine sensitivity and specificity of the assay. This project uses site-directed mutagenesis to generate a panel of BRCA2 variants for functional assay development and validation.
Introduction
Methods
ConclusionsOf the 16 BRCA2 variants made across two semesters, 13 were confirmed with sequencing, 1 failed at the transformation step, and 2 could not be confirmed with sequencing. Four variants that didn’t form colonies in the first semester, were successful and confirmed with sequencing when made during the second semester.These initial failed attempts could be attributed to not being incubated at the correct temperature during transformation due to a broken orbital shaker.The PCR protocol was also modified to use 25 ng of starting DNA rather than the original 10 ng, and the denaturation time was maximized to increase transformation efficiency (see Figure 7).Future Directions?Assess expression at protein level (Western blots)Test functional assay (traffic light reporter assay) – can the assay distinguish between the benign and pathogenic variants in the validation panel?Analyze BRCA2 VUS with functional assay – can it help reclassify some of the VUS?Analyze PALB2 VUSChallengesTime to completion is a barrier to high-throughput adaptationCost of sequencing to confirm successful mutagenesisDifficult to make multiple changes in the gene at once
References:Breastcancer.org. U.S. Breast Cancer Statistics. Ardmore, PA. Breastcancer.org; 9 Jan 2018.Guidugli, L., Carreira, A., Caputo, S. M., Ehlen, A., Galli, A., Monteiro, A. N.A., Neuhausen, S. L., Hansen, T. V.O., Couch, F. J., Vreeswijk, M. P.G. and on behalf of the ENIGMA consortium (2014), Functional Assays for Analysis of Variants of Uncertain Significance in BRCA2. Human Mutation, 35: 151–164. DOI:10.1002/humu.22478Richards, S., Aziz, N., Bale, S., Bick, D., Das, S., Gastier-Foster, J., … Rehm, H. L. (2015). Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genetics In Medicine, 17, 405. Retrieved from http://dx.doi.org/10.1038/gim.2015.30 Laursen, Kristian. Site Directed Mutagenesis by PCR. Addgene; 2 Aug. 2016. QuikChange II XL Site-Directed Mutagenesis Kit. Agilent Technologies, 2015. Bellcross, Cecelia. The Genetics of Early-Onset Breast Cancer (PowerPoint). Department of Human Genetics. Emory University School of Medicinee. Atlanta, GA.ClinVar Database. https://www.ncbi.nlm.nih.gov/clinvar/. Accessed April 2018.
Acknowledgements:
This work was supported in part thanks to the
UNC School of Medicine Yang Family Biomedical Scholars Award.
What is Site-Directed Mutagenesis?PCR-based approach to making small, targeted changes in DNA.4Primers are complementary to the gene of interest with the exception of a single, specific point mutation, and PCR introduces the change in resulting amplified DNA (Figure 6).The parental DNA is enzymatically digested, leaving only DNA containing the mutation.DNA is transformed into competent bacterial cells and colonies are grown on selection media.Colonies are isolated and grown in cultures overnight.The DNA is extracted and Sanger sequencing confirms the successful incorporation of the specific mutation.Why use Site-Directed Mutagenesis?High-fidelity mutagenesis: highly specific, few off-target resultsGood choice for mutagenesis of plasmid DNA
Evidence
types used to classify variants
3
:In silico (computational) predictive methodsSegregation studiesFunctional assaysEven in genes with established roles in disease (e.g. BRCA2 and breast cancer), VUS are numerous and problematic (Figure 2).12% of U.S. women will develop breast cancer in their lifetime (Figure 3). 5-10% of these cases are due to a hereditary syndrome.1Genes associated with hereditary breast cancer normally act to repair DNA double-strand breaks by homologous recombination (HR).BRCA1BRCA2PALB2Identifying pathogenic germline variants in these genes is clinically important. Risk managementPatient counselingTreatment decisionsFunctional assay development is important for variant classification.The ENIGMA consortium has published a panel of “genetically proven pathogenic and neutral [benign] missense variants” for BRCA2 assay validation.2
The purpose of this research is…to implement site-directed mutagenesis to generate a validation panel of previously classified BRCA2 variants that can be used in the development of a robust functional assay of HR pathway germline variants that will aid in clinical pathogenicity determinations.
Figure 8. PCR Optimization
Figure 9. Examples of successful transformation
Figure 7.
QuikChange Method Overview5
Figure 2
Benign
Likely Benign
VUS
Likely Pathogenic
Pathogenic
BRCA2 Protein Functional Domains
(
Mutagenesis Target Sites Indicated
by Arrows)
N-terminus
BRCA2 Repeats (RAD51)
DNA Binding Domains
HD
OB1
OB2
OB3
C-terminus
R18H
L109V
F1524V
H2074N
K2472T
T2515I
I2627F
L2653P
L2647P
T2722R
L2688P
D2723G
K2729N
N3124I
G2748D
Figure 3
Figure 6.
Sanger confirmation
BRCA2 I2627F (
c. 7879 A>T)
PCR was performed as part of the mutagenesis of the BRCA2 protein mutation I2674F (2/15/18).
Number
of Colonies
Controls
K2472T
L2688P
G2748D
L2653P
>100
(PCR control)
13
27
43
73
33 (transformation control)
14
64
56
77
Figure 5. Work Flow Overview
Figure 1
. American College of Medical Genetics and Genomics (ACMG) Guidelines for Clinical Variant Interpretation.
3