PPT-Details of polymer chain
Author : karlyn-bohler | Published Date : 2019-06-20
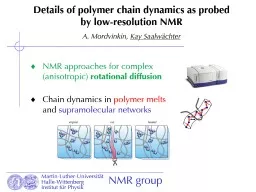
dynamics as probed by lowresolution NMR A Mordvinkin Kay Saalwächter NMR approaches for complex anisotropic rotational diffusion Chain dynamics in polymer
Presentation Embed Code
Download Presentation
Download Presentation The PPT/PDF document "Details of polymer chain" is the property of its rightful owner. Permission is granted to download and print the materials on this website for personal, non-commercial use only, and to display it on your personal computer provided you do not modify the materials and that you retain all copyright notices contained in the materials. By downloading content from our website, you accept the terms of this agreement.
Details of polymer chain: Transcript
dynamics as probed by lowresolution NMR A Mordvinkin Kay Saalwächter NMR approaches for complex anisotropic rotational diffusion Chain dynamics in polymer melts and . Presented by. Steve Albrecht. Technical Director. Adherent Laboratories. 50 Years and Counting…... Early 1970’s – Technology was EVA/. tackifying. resin blends. Low cohesive strength. Not good performance at body temperature. for the Treatment of . Oral and Head And Neck Carcinoma. MAIE A. ST. JOHN, MD, PHD. Department . of Head & Neck Surgery. David Geffen School of Medicine, UCLA. Jonsson Comprehensive Cancer . Center. Nanoparticle Fabrication via . Photodimerization. of P. endant Anthracene ROMP . Polymer. Mark F. Cashman, Peter G. Frank*, Erik B. . Berda. *. Department of Chemistry, University of New Hampshire, Durham, NH. . DEFINITION . A Polymer is a substance composed of molecules with a large molecular mass of repeating structural units or monomers connected by covalent chemical bonds.. . Cross . linked Polymer:. . A lesson on Polymers. Outline. Basics of polymers. What is a polymer?. What is a thermoplastic?. What is a thermoset?. Unusual Behavior of Polymers (Thermoplastics). Weissenberg. Kaye. Barus. Anti-Gravity. Sash Chain. Sash Chain is sold by the foot. Used to hang light fixtures, etc. . Twist Link Chain. Used where the chain must travel easily over something (links don’t get caught).. Double Loop Chain. Fei. Yu and . Vikram. . Kuppa. School of Energy, Environmental, Biological and Medical Engineering. College of Engineering and Applied Science. University of Cincinnati. APS . March . Meeting 2012, Boston. Super Absorbent Polymer Market report published by Value Market Research is an in-depth analysis of the market covering its size, share, value, growth and current trends for the period of 2018-2025 based on the historical data. This research report delivers recent developments of major manufacturers with their respective market share. In addition, it also delivers detailed analysis of regional and country market. View More @ https://www.valuemarketresearch.com/report/super-absorbent-polymer-market 2 MATERIAL21 Polymer synthesisThe polymer studied is based on the structure shown in Fig 1 whichis called hereafter 3RDCVXY It is now wellunderstood that for an organic molecule to possess large secon structural isomerism is used when polymers have the same overall chemical composition (i.e., same molecular formula) but differ in . the . order in which the atoms are connected to each other. . Polyacetaldehyde. By. Asst. Prof. Dr. Raouf Mahmood Raouf. Introduction. Clay, a natural source, with small loadings (%by weight) can substitute . reinforces . which are being used in polymers. The commercial importance of polymers has led to an strong investigation of polymeric material nanocomposites, of sizes varying from . Bioresorbable. Polymer-Based Drug-Eluting Stent in an All-Comers Patient Population. Lisette Okkels Jensen, Michael . Maeng. , Bent . Raungaard. , Johnny . Kahlert. , Julia Ellert, Lars . Jakobsen. , . Katerina Lazarova. 1*. , Rosen Georgiev. 1. , . Darinka. Christova. 2. and . Tsvetanka. Babeva. 1*. 1. Institute . of Optical Materials and Technologies ‘‘Acad. J. Malinowski’’, Bulgarian Academy of Sciences, Acad. G. . : The. . HOST. -Reduce-. Polytech. -. ACS. . trial. Hyo-Soo Kim, MD/PhD. Cardiovascular Center,. Seoul National University Hospital, Korea. Session of Late Breaking Trial Session IV. Co-sponsored by European Heart Journal.
Download Rules Of Document
"Details of polymer chain"The content belongs to its owner. You may download and print it for personal use, without modification, and keep all copyright notices. By downloading, you agree to these terms.
Related Documents