PPT-Let’s
SO
mitsue-stanley
Published 2018-01-07 | 5144 Views
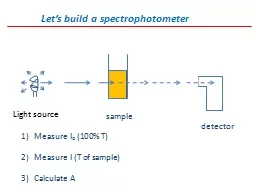
build a spectrophotometer Light source sample detector Measure I o 100 T Measure I T of sample Calculate A Change the monchromator to measure A versus wavelength
Download Presentation
Download Presentation The PPT/PDF document "Let’s" is the property of its rightful owner. Permission is granted to download and print the materials on this website for personal, non-commercial use only, and to display it on your personal computer provided you do not modify the materials and that you retain all copyright notices contained in the materials. By downloading content from our website, you accept the terms of this agreement.
