PPT-Mapping the Free Energy Landscape of HIV-1 TAR RNA with
Author : phoebe-click | Published Date : 2017-09-04
Metadynamics Tyler J Mulligan 1 Harish Vashisth 2 1 MS Student Department of Chemical Engineering University of New Hampshire Durham NH 2 Advisor Department of
Presentation Embed Code
Download Presentation
Download Presentation The PPT/PDF document "Mapping the Free Energy Landscape of HIV..." is the property of its rightful owner. Permission is granted to download and print the materials on this website for personal, non-commercial use only, and to display it on your personal computer provided you do not modify the materials and that you retain all copyright notices contained in the materials. By downloading content from our website, you accept the terms of this agreement.
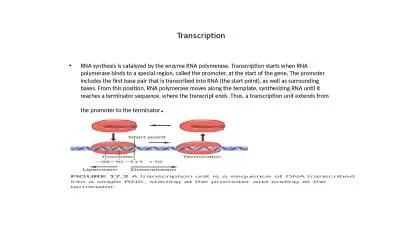
Mapping the Free Energy Landscape of HIV-1 TAR RNA with: Transcript
Download Rules Of Document
"Mapping the Free Energy Landscape of HIV-1 TAR RNA with"The content belongs to its owner. You may download and print it for personal use, without modification, and keep all copyright notices. By downloading, you agree to these terms.
Related Documents