PPT-A+ C+
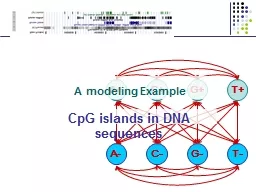
G T A C G T A modeling Example CpG islands in DNA sequences Methylation amp Silencing One way cells differentiate is methylation Addition of CH 3 in Cnucleotides
Download Presentation
"A+ C+" is the property of its rightful owner. Permission is granted to download and print materials on this website for personal, non-commercial use only, provided you retain all copyright notices. By downloading content from our website, you accept the terms of this agreement.
Presentation Transcript
Transcript not available.