PPT-Recombination breakpoints
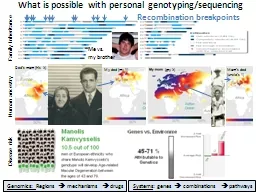
Family Inheritance Me vs my brother My dad my Y Moms dad uncles Y Human ancestry Disease risk Genomics Regions mechanisms drugs Systems genes combinations pathways
Download Presentation
"Recombination breakpoints" is the property of its rightful owner. Permission is granted to download and print materials on this website for personal, non-commercial use only, provided you retain all copyright notices. By downloading content from our website, you accept the terms of this agreement.
Presentation Transcript
Transcript not available.