PPT-Random Genetic Drift
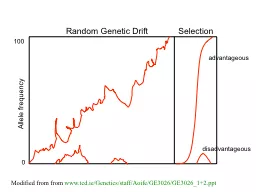
Selection Allele frequency 0 100 advantageous disadvantageous Modified from from wwwtcdieGeneticsstaffAoifeGE3026GE302612ppt Purifying selection in GTA genes dNdS
Download Presentation
"Random Genetic Drift" is the property of its rightful owner. Permission is granted to download and print materials on this website for personal, non-commercial use only, provided you retain all copyright notices. By downloading content from our website, you accept the terms of this agreement.
Presentation Transcript
Transcript not available.